Summary
Phenotypic plasticity allows organisms to alter gene expression and phenotype in response to environmental changes. Recent evidence highlights environmentally induced changes in gene regulation, revealing the environment as both a selective agent and a phenotypic inducer. Genetic accommodation, where the degree of plasticity is modulated in response to persistent environmental conditions, suggests an evolutionary mechanism driven by plasticity, though its underlying mechanisms remain largely unknown. To fully understand the mechanisms behind phenotypic plasticity and genetic accommodation, exploring these processes at finer geographical and temporal scales than speciation is crucial. We investigated adaptive developmental plasticity in Pelobates cultripes, a highly plastic spadefoot toad species spread in the Iberian peninsula, responsive to environmental stressors like pond drying. Using a transcriptomic RNA-seq approach, we compared gene expression profiles of P. cultripes tadpoles from Central and Southern Spain populations exposed to different water levels. Here, we seek to identify differentially expressed genes (DEGs) involved in these tadpoles larval development acceleration under desiccation stress. Our results indicate that Southern Spain populations exhibit a higher number of DEGs in response to low water levels compared to Central Spain populations, possibly accounting for the different plasticity levels of populations and developmental rates. By examining gene expression patterns across diverse populations and environmental conditions, we underscore the importance of comprehensive studies on genetic accommodation, contributing to a broader understanding of how organisms adapt to rapidly changing environments.
The RNAseq Workflow
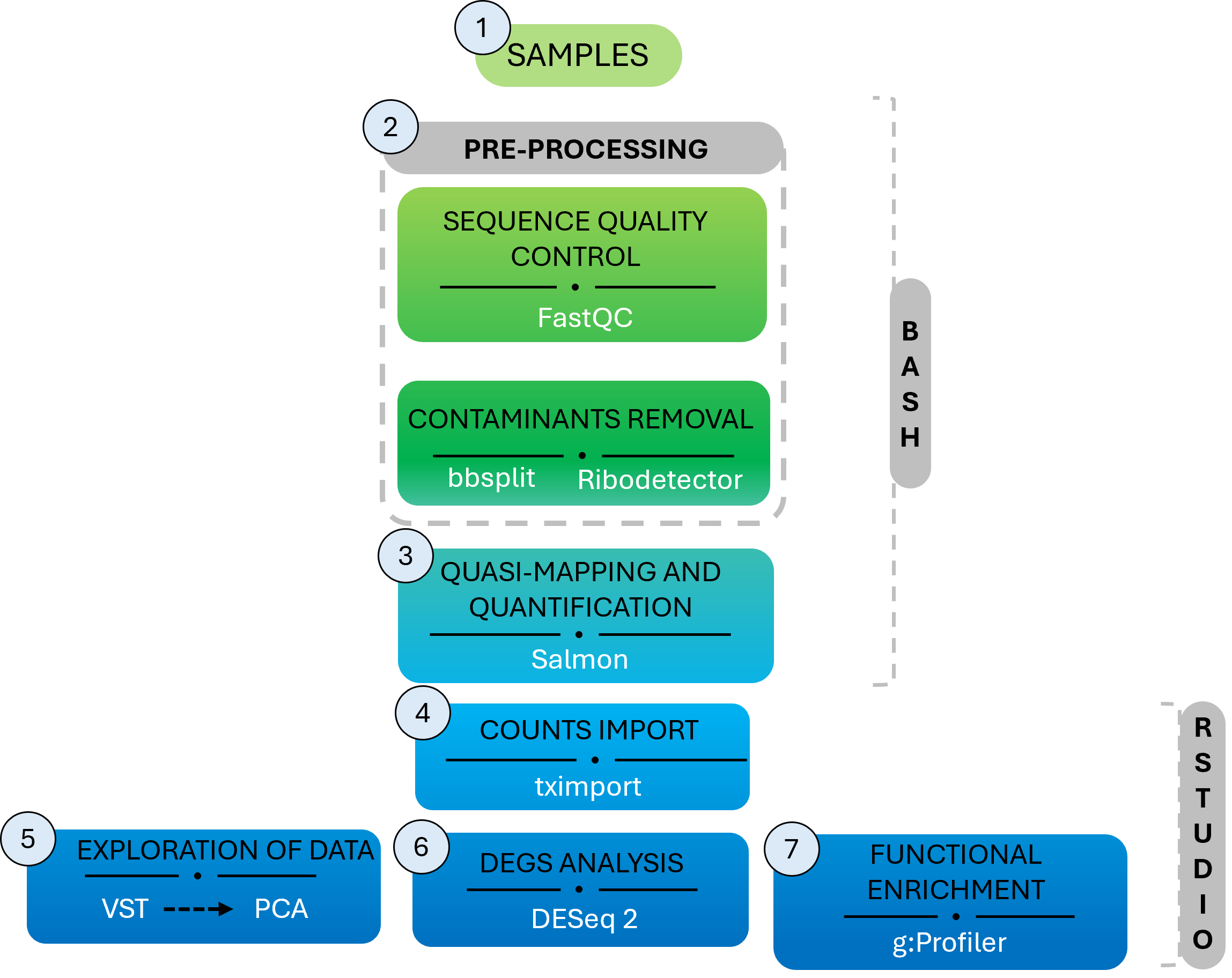
-
Pre-processing
-
Differential Expression Analysis
About The Research Group
Learn more about the research group in the following web Eco-Evo-Devo Lab.